Infection Models
Immortalised cell lines are frequently used in infection research, but their applications are restricted particularly when immune responses of humans or animals to a viral infection are investigated.
The immune response is a complex interaction of various components of the immune system, and up to now it cannot be mimicked completely in cell culture systems. Furthermore, the diversity of cell types gets lost in immortalised cell lines and changes in expression patterns of cellular proteins may occur. Cells of the respiratory tract are particularly subject to the loss of typical characteristics, such as the production of mucus or the presence of cilia. Thus, cell cultures cannot or do not sufficiently reflect the situation of organs in vivo. For more in-depth analyses of interactions between pathogens and hosts, it is inevitable to use models that ideally copy the situation within the living organism.
The main aim of our platform is to establish and to continuously develop various models while steadily considering the 4R principles (replacement, reduction, refinement, responsibility). Our models include both in vivo models in non-human primates (NHP) and ex vivo models using primary cell cultures.

Primary cell cultures reflect the in vivo situation of selected organs very well; depending on the culture, the organ-typical cell association is completely preserved. In contrast to immortalised cells, the cells are unchanged and continue to possess their specific physiology and function. As primary cell cultures may be gained from organ samples, e.g. biopsy material, or from recently deceased organisms, there is no need to use laboratory animals to generate these cultures.
In vivo models have the highest complexity and are especially suitable for research questions concerning transmission, pathogenesis and immune response of infectious diseases. For ethical and animal welfare reasons, in vivo models may only be applied if the scientific results (i) is not sufficiently understood, (ii) show a benefit for the health of human and/or animal, and (iii) cannot not be obtained from another less complex model.
NHP models
The close evolutionary relationship between humans and primates makes them suitable models to examine and to comprehend different aspects of infectious diseases in humans. This implies transmission paths, pathogenesis and therapeutic approaches such as the use of monoclonal antibodies against specific viral infections or the testing of new immunization strategies and vaccines for antiviral prophylaxis.The choice of primate models depends on the specific issue and the virus to be analysed. Rhesus monkeys, long-tailed macaques and marmosets are among the most frequently used animal models. Our facilities enable us to pursue and evaluate infection trials with biological and genetically modified pathogens of biosafety levels ranging from 1 to 3 in accordance with the effective Biological Agents Ordinance, the Specified Animal Pathogens Ordinance and the Genetic Engineering Act.Our already established models in rhesus and long-tailed macaques include for example an SIV/HIV (simian/human immunodeficiency virus) primate model to analyze HIV, models to investigate novel therapeutical options with adeno-associated viruses (AAV) as well as infection models on various respiratory viruses, such as the respiratory syncytial virus (RSV), influenza A viruses (FluA) and SARS-CoV-2. Furthermore, a marmoset model has been established for research on the Kaposi sarcoma.
Routine sampling is mainly done via collection of blood samples, swabs samples, urine samples, bronchoalveolar lavage fluid and, depending on the circumstances and the pathogen, via bone marrow puncture, collection of cerebrospinal fluid and lymph node biopsies under injection anaesthesia. Major surgical interventions are done by inhalation anaesthesia and continuous anaesthesia monitoring.
Primary cell cultures
In certain cases, primary cell cultures are an alternative in terms of animal models and contribute to the reduction of the number of experimental animals according to the 4R principle. Our focus lays on primary cultures of the respiratory tract. Especially precision-cut lung slices (PCLS) represent an ex vivo model which enables us to examine the susceptibility of lung cells to certain respiratory pathogens, to identify target cells and to test antiviral agents in a setting which reflects lifelike lung conditions.
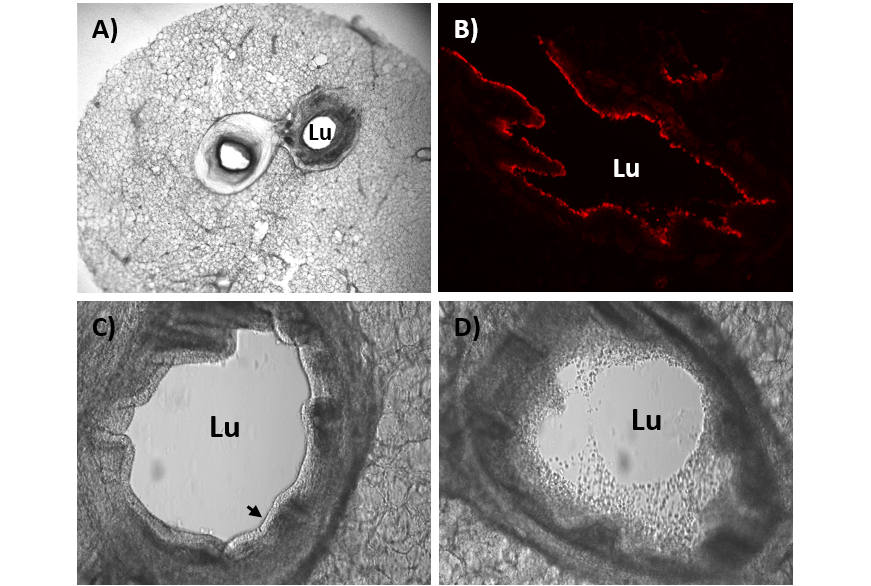
B) The lumen (Lu) of the bronchi is surrounded by ciliated epithelial cells (stained red) and mucus producing cells. Mucus and cilia are important defense mechanisms of the respiratory tract and protect against foreign bodies e.g. dust and infectious agents.
C) The ciliated epithelium (see arrow) lines the bronchus of a healthy lung.
D) Following a viral infection, dead cells of the ciliated epithelium can be found in the lumen of the bronchus. The local defense mechanism of the lung is impaired by the cell death.
Selected publications:
Färber I, Krüger J, Rocha C, Armando F, von Köckritz-Blickwede M, Pöhlmann S, Braun A, Baumgärtner W, Runft S, Krüger N. „Investigations on SARS-CoV-2 Susceptibility of Domestic and Wild Animals Using Primary Cell Culture Models Derived from the Upper and Lower Respiratory Tract.” Viruses. 2022 Apr 16;14(4):828. doi: 10.3390/v14040828.
Krüger N, Rocha C, Runft S, Krüger J, Färber I, Armando F, Leitzen E, Brogden G, Gerold G, Pöhlmann S, Hoffmann M, Baumgärtner W. „The Upper Respiratory Tract of Felids Is Highly Susceptible to SARS-CoV-2 Infection.” Int J Mol Sci. 2021 Sep 30;22(19):10636. doi: 10.3390/ijms221910636.
Hempel T, Elez K, Krüger N, Raich L, Shrimp JH, Danov O, Jonigk D, Braun A, Shen M, Hall MD, Pöhlmann S, Hoffmann M, Noé F. „Synergistic inhibition of SARS-CoV-2 cell entry by otamixaban and covalent protease inhibitors: pre-clinical assessment of pharmacological and molecular properties.” Chem Sci. 2021 Aug 26;12(38):12600-12609. doi: 10.1039/d1sc01494c.
